Last year's Schol Bio paper contained (as is usual) some interesting and challenging questions. One of them was about earwax. More specifically, the earwax phenotypes 'dry' and 'wet', and what their distribution can tell us about patterns of human evolution. (Note to those sitting these examinations: most questions have a reasonable amount of resource material provided and this one was no exception. Remember to use this information in your answers! Ignoring it, or – just as bad – copying bits rather than incorporating the material properly – is not a sign of a good response.)
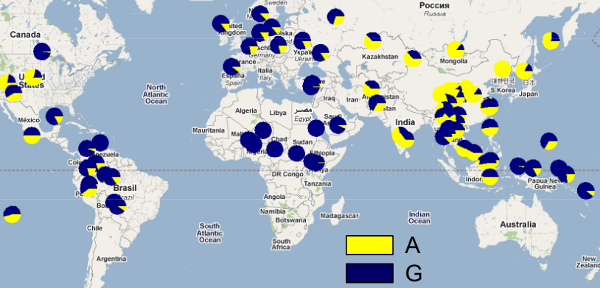
I was particularly interested in this question because we used to look at the distribution of various phenotypes, wet & dry earwax among them. The examiner provided information about the nature of the genetic information underpinning the two phenotypes, along with data about the global distribution of people with wet & dry wax in their ear canals, including a very helpful map. The following graphic is not that map, but is very similar – showing allele frequencies rather than the phenotype distributions used in the exam – & accompanies an excellent blog post on the Discover magazine site. The 'A' and 'G' represent the 2 alleles involved in determining the relative dampness of your earwax:
Earwax is wet unless an individual has Adenine (A) at a particular site instead of Guanine (G), in which case the wax becomes the dry form. People who inherit the version of the gene that has A from both parents have dry earwax. People who inherit two of the G versions, or one G and one A, have wet earwax.
The actual question asked candidates to
use biological knowledge, together with information from the resource material, to discuss:
- the origins and inheritance patterns of dry earwax
- the evolutionary factors that may have resulted in the present-day distribution of both types of earwax.
And as always, a successful candidate would address all parts of a question. Nor would they assume that the examiner 'knows' what they know – you do need to spell out your understanding.
So, for the first bullet point: we're dealing with a substitution mutation, where a change in a single base (from G to A) has led to a single amino acid change in the final protein. That secretory protein's function is altered so that the wax that's produced is now 'dry'. The mutation must have occurred in a gamete-producing cell, or at the least during meiosis, & subsequently entered the human population's gene pool. We know it's a recessive mutation as someone must be homozygous for the allele to express wet wax, while heterozygotes have dry earwax. And you could also add that it's not a sex-linked mutation, because (as the resource material notes) the gene involved in wax secretion is found on chromosome 16.
The second part of the question requires you to relate the information on the distribution of 'dry' & 'wet' wax phenotypes to your knowledge of patterns of human dispersal (in this case, the 'out-of-Africa' model). The fact that there's no 'A' allele in African populations suggests that this mutation must have arisen after our species started to spread out of Africa. Furthermore, it could well have appeared once a small founder population had arrived in China, with subsequent genetic drift removing the dominant (G) allele from that founder group – this would explain the very high frequency of the A allele (up to 100%) in that region. You could also argue the possibility of positive selection pressure on this version of the secretory protein (an idea critiqued by the author of the Discover blog post).
However, the A allele is also found at fairly high frequencies in other Asian countries – the most likely explanation here is that subsequent migration and interbreeding has introduced it to those populations (with 54% of Indians and 69% of Japanese now expressing the 'dry' phenotype). Until fairly recently there was only minimal migration from China into Russia & Europe, which accounts for the very low frequency of the recessive allele in those populations.
What about the Americas? Our current understanding is that humans migrated into North America from Asia via a land bridge across the Bering Strait, during a glacial period. (There's an interesting discussion around this here, including some work done using a molecular clock based on mutations in the common human pathogen Helicobacter pylori to estimate migration paths & times.) In dealing with this part of the question, I can think of a couple of options: that the lower frequency of the A allele in native American peoples reflects a founder event where the allele was at lower frequency to begin with, & subsequent genetic drift; or that the A allele frequency originally reflected that in Asian populations, but was diluted by a significant drop in population size following European settlement & some later interbreeding with the settlers. Simiilarly, the low A frequency in non-native Americans simply reflects their relatively recent arrival from Europe.
Who'd have thought that the story of earwax could be so fascinating?

herr doktor bimler says:
some later interbreeding with the settlers
“Settlers” includes the involuntary immigrants from Africa who (I suspect) contributed to the high proportion of the G allele in North and South America — higher than European immigrants could explain, for some of those data points.
Andrew Broome says:
http://www.sott.net/article/267961-Ancient-DNA-links-Native-Americans-with-Europe
Jim Thomerson says:
Is it correct to think that distribution of phenotypes pretty closely follows Hardy-Weinberg?
Alison Campbell says:
HW looks at allele and genotype frequencies in a particular population at a particular point in time; if those frequencies have changed at a later sampling time then one could assume there was some evolutionary pressure on the gene pool. But I don’t think it’s accurate to say that HW can be applied to the global population.